Bacteria secrete an incredible array of chemicals that affect their environment, neighboring microbes, and their hosts. But only a small number of these chemicals have been identified and characterized.
To improve detection of secreted microbial chemicals, Pieter Dorrestein of the University of California, San Diego, and colleagues employed a recently developed mass spectrometry (MS) technique called nanoDESI (nanospray desorption electrospray ionization), which enables the repeated sampling of living microbes right from petri dishes.
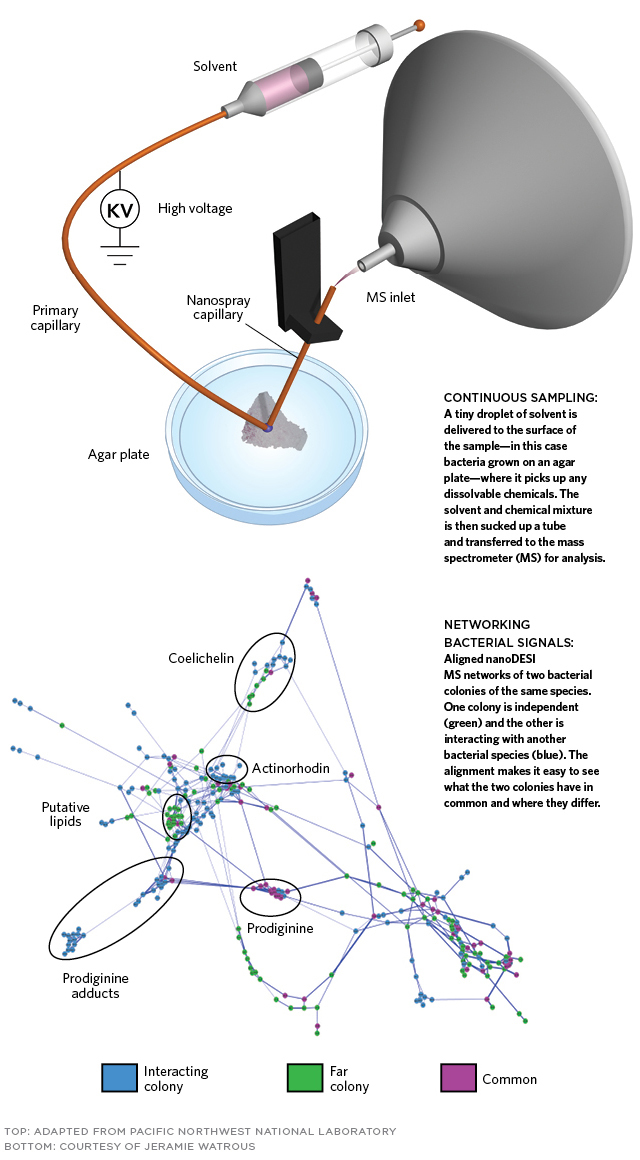
In nanoDESI a tiny drop of solvent—approximately 0.2 millimeter in diameter—is delivered to the surface of the target sample and then sucked slowly by capillary action into another tube along with any chemicals picked up from the sample cells. The solution is then analyzed by MS. In this case, the sample was a bacterial colony; it was the first time nanoDESI had been used to repeatedly sample from living bacteria.
The approach generated an unprecedented amount...
The software has enabled the identification of a number of new microbial products, including one called thanamycin, a lipopeptide secreted by Pseudomonas bacteria in the soil that protects sugar beet plants from fungal infections. The chemical was predicted to exist, but had long evaded detection.
“Professor Dorrestein’s study constitutes a paradigm shift for microbiology,” says Catherine Fenselau, a biochemist at the University of Maryland, who says the approach will be readily adapted by others. (PNAS, 109:E1743–52, 2012.)
.
| STATS TALK | |||||
| THE METHODS: | HOW IT WORKS | ADVANTAGES | DISADVANTAGES | ||
| LC-MS | Liquid chromatography-MS followed by database searching | A number of computer programs are available for comparing MS signatures. | Limited number of annotated molecules in databases makes identification of molecules difficult. | ||
| NanoDESI-MS with Dorrestein’s software | NanoDESI-MS followed by alignment of spectral networks | Aligning networks makes differences easy to observe. Used together with database searching, it increases chances of classifying unknown molecules. | Less abundant metabolites are difficult to detect. | ||
Interested in reading more?





