 MARIA AND PETER HOEY
MARIA AND PETER HOEY
Cells live longer than their internal components. To keep their cytoplasm clear of excess or damaged organelles, as well as invading pathogens, or to feed themselves in time of nutrient deprivation, cells degrade these unwanted or potentially harmful structures, and produce needed food and fuel, using a process they have honed over millions of years. Known as autophagy, this catabolic process involves the selection and the sequestration of the targeted structures into unique transport vesicles called autophagosomes, which then deliver the contents to lysosomes where they are degraded by lytic enzymes. This conserved eukaryotic pathway plays a central role in a multitude of physiological processes, including programmed cell death, development, and differentiation. In addition, it plays a protective role against aging, tumorigenesis, neurodegeneration, and infection. Given all this, it is not surprising that an impairment of autophagy is correlated with various severe pathologies, including cardiovascular...
Despite significant advances over the last 20 years in the understanding of how this process works and what purposes it serves, there is a lingering question—how are autophagosomes formed? More specifically, where do their not one, but two lipid bilayers come from? Autophagosomes are not pre-built organelles that become active upon the induction of autophagy; they are made from scratch each time a cell needs to degrade one or more of its contents. And they are giant vesicles, with an average diameter of approximately 700–800 nanometers, which can further expand to accommodate large structures such as cellular organelles and bacteria, and which are made in large quantities under autophagy-inducing conditions. As a result, progression of autophagy requires a ready supply of lipids. This aspect of the process has intrigued researchers since the discovery of autophagy in the 1950s and ’60s. Understanding the biogenesis of autophagosomes will provide information about how cells generate new compartments in response to internal and external cues, and will thus lead to a clearer conception of cell homeostasis.
Intrinsic to the question of the autophagosome’s origin is the source of the lipids required to build the double-membrane vesicle and the way this supply is delivered. One major difficulty in addressing this question has been that autophagosomes contain no marker proteins that definitively link them to any known subcellular organelle, making it difficult to unveil their origins. Indeed, autophagosomes are distinct from all other organelles in the cell, both in structure and in protein composition. Recent advances in microscopic techniques and biochemical approaches have stimulated a series of studies investigating this issue, but the results are contradictory, at least at first glance, with different groups identifying evidence for contributions from the cell’s plasma membrane, endoplasmic reticulum, mitochondria, and Golgi complex. From which of these organelles is the autophagosome derived, or could it be all of the above? The answer to this question is a prerequisite for understanding and manipulating the mechanism of autophagy. In turn, this knowledge is essential to the development of therapies or drugs that target this pathway to treat or even cure diseases in which autophagy is blocked or impaired.
How autophagy works
Autophagosome biogenesis and consumption can be divided into five discrete steps: induction, expansion, vesicle completion, fusion, and cargo degradation. The initial event upon induction is the formation of a membranous cistern called the phagophore, or isolation membrane. This compartment appears to be generated from what has been defined in yeast as the phagophore assembly site (PAS) or pre-autophagosomal structure, a putative early autophagosome precursor that is formed by the sequential association of at least a subset of Atg proteins, which are known to be specifically involved in autophagy. The subsequent expansion of the phagophore through the acquisition of additional lipids permits the engulfment of the intracellular material targeted for destruction. The double-membrane vesicle is completed when the inner and outer bilayers fuse to form two distinct membranes, one inside the other. Completed autophagosomes first fuse with endosomal structures to form amphisomes (an event that appears not to occur in yeast), and then with the mammalian lysosome or the yeast and plant vacuole, allowing the degradation of the inner vesicle and its cargo by acid hydrolases residing in these lytic compartments. Lastly, the basic metabolites generated from this catabolic processing of biological macromolecules are transported into the cytoplasm, where they are reused as either a source of energy or as building blocks for new proteins and lipids.
Mixed membrane messages
THE ENDOPLASMIC RETICULUM
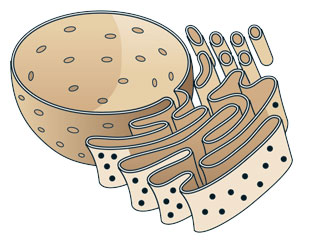 Endoplasmic reticulumMARIA AND PETER HOEY
Endoplasmic reticulumMARIA AND PETER HOEY The first organelle proposed as the source of autophagosomal membranes was the endoplasmic reticulum (ER), the compartment responsible for the production of the proteins and lipids that compose the cell. Morphological studies performed in the 1970s already indicated a possible functional link between autophagosomes and the ER, because these two organelles were often seen in close proximity. These observations were subsequently supported by immuno-ultrastructural analyses made by Bill Dunn of the University of Florida in 1990, in which integral membrane proteins of the rough ER were detected in both the inner and outer membranes of the autophagosomes.
More recently, using electron microscope tomography to study the three-dimensional organization of these organelles, two groups have confirmed and emphasized these pioneering observations by revealing the existence of a physical connection between the ER and the forming autophagosomes. In 2009, Mitsuko Hayashi-Nishino and colleagues at Osaka University in Japan performed 3-D reconstruction of cells expressing a mutant gene that causes defects in autophagosome formation, effectively pausing the process at the early stages of autophagosomal membrane formation (i.e., the phagophore stage).1 This analysis revealed that the ER and the growing phagophore are intimately associated, suggesting that the nascent autophagosome branches off from the ER. In fact, the rough ER was connected through a single point of contact to both the outer and inner membranes of the phagophore, supporting the notion that lipids could be supplied via direct transfer at the sites of membrane contact. It is unclear, however, whether phagophore membrane formation starts at the ER, or whether the membrane simply grows there after the process is initiated elsewhere. Päivi Ylä-Anttila of the University of Helsinki and co-workers obtained similar findings in a different cell type, but they have reported the existence of several points of contact between these two organelles.2
Further support for the involvement of the ER in autophagosome formation comes from studying phosphatidylinositol-3-phosphate [PI(3)P], a lipid crucial for autophagosome formation. In 2008, researchers found that PI(3)P is enriched in specific regions of the ER where the autophagosomes have been observed breaking off under autophagy-inducing conditions.3 The same group also showed that cup-shaped structures they called omegasomes, which may be an autophagosome precursor (probably a phagophore expansion intermediate), emerge from these PI(3)P-rich subdomains. These findings have been corroborated by a new report revealing that Atg14L, a subunit of the complex involved in the synthesis of PI(3)P, is associated with the ER surface.4 Similarly, DFCP1, a protein mainly localizing to the ER, was found to be associated with both omegasomes2 and autophagosomal membranes connected to the ER.1
Taken together, these data suggest that autophagosomes emerge from the ER. However, 30 percent of the autophagosome precursors observed in the 3-D tomography studies were not associated with the ER, raising the possibility that another membrane source for autophagosomes could exist.1
THE MITOCHONDRION
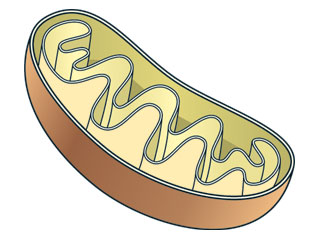 MitochondrionMARIA AND PETER HOEY
MitochondrionMARIA AND PETER HOEY In 2010, Jennifer Lippincott-Schwartz’s group at the National Institutes of Health in Bethesda proposed the outer membrane of the mitochondria as the main source of the autophagosomal lipid bilayers.5 Their findings were obtained from extensive light-microscopy analyses of cells expressing fluorescently tagged marker proteins for autophagosomes and proteins localizing to the mitochondrial outer membrane. Under amino acid starvation conditions that initiate autophagy, these two sets of marker proteins were found to co-localize on nascent autophagosomes, suggesting a functional link between mitochondria and autophagosomes. The researchers also observed a potential direct physical connection between these two organelles, with autophagosomes growing in close proximity to mitochondria, leading the authors to propose that the mitochondria supply the forming autophagosomes with newly synthesized phospholipids.6,7,8 More recently, however, another group found that Salmonella-containing autophagosomal structures were negative for the same mitochondrial marker protein, indicating that mitochondrial lipids may not be involved in the biogenesis of all autophagosomes.9
Lippincott-Schwartz’s team also showed that the connection between the mitochondria and the ER is crucial for autophagosome formation. Mitochondria are normally associated with the ER through discrete points of contact that are known as the mitochondrial-associated membranes (MAMs). In the absence of MAMs, phospholipid biosynthesis is impaired, indicating their important role in the lipid exchange between ER and mitochondria. When MAMs are disrupted by knocking down one of the genes involved in their maintenance, starvation-induced autophagosomes are not formed.5 This observation has led to the hypothesis that the ER contribution as a membrane provider for autophagy could be as important as the mitochondrial one. Another possible interpretation, however, is that loss of the connection between the two organelles could lead to an impairment of several functions of the ER, which in turn could affect its contribution to phagophore or autophagosome biogenesis and/or expansion.
THE PLASMA MEMBRANE
Yet another possible contributor to the autophagosomal double membrane is the cell’s plasma membrane. In 2010, the laboratory of David Rubinsztein at the University of Cambridge reported that some vesicles forming from the plasma membrane are positive for the autophagosome marker Atg16L1, and fuse with like vesicles to create an early autophagosomal precursor, possibly a phagophore or an omegasome.10 Additional investigations from the same group have revealed that the maturation of Atg16L1-positive pre-autophagosomal vesicles requires their fusion via the action of a plasma membrane SNARE protein and its interacting partners. The involvement of plasma membrane SNAREs, which mediate vesicle fusion and exocytosis, in the early steps of autophagy has also been highlighted by studies in yeast. Additional support for a plasma membrane role in autophagosome biogenesis comes from the discovery that components of the exocyst—a tethering complex that acts in concert with SNAREs to mediate fusion of Golgi-derived vesicles with the plasma membrane—associate with nascent autophagosomes, and that they are essential for starvation-induced autophagy.
THE GOLGI APPARATUS
 Golgi ApparatusMARIA AND PETER HOEY
Golgi ApparatusMARIA AND PETER HOEY The potential involvement of the exocyst in autophagosome formation also raises the possibility of the Golgi apparatus being a potential source for autophagosomal membranes, as the exocyst complex is present in the Golgi as well as in vesicles derived from them. The Golgi is an organelle dedicated to posttranslational protein modifications and sorting. The work on yeast plasma membrane SNAREs has revealed that these proteins regulate the organization of organelles containing Atg9, a transmembrane protein essential for autophagy. In yeast, these organelles, which have been named Atg9 reservoirs, appear to be derived from the Golgi and play a central role in providing the initial membranes necessary to generate the phagophore.11 Even as early as the 1990s, the Golgi was implicated in autophagosome biogenesis in mammalian cells, as the growing phagophore, as well as complete autophagosomes, could be decorated with proteins that bind to sugar chains exclusively present on post-Golgi membranes. This concept has been recently reinforced by work in yeast, where it has also been shown that two complexes involved in membrane trafficking through the Golgi are essential for autophagy.
A single model?
While researchers have accumulated undoubtedly confusing evidence for the involvement of multiple organelles in the formation of autophagosomes, the resulting hypotheses are not mutually exclusive. Further research is required to sort out which organelles contribute to autophagosome biogenesis in which species under which conditions.
The apparent discrepancy between the conclusions reached by different laboratories on the origin of the autophagosomal membrane could be, in part, due to varying experimental approaches and techniques. More importantly, the different contributions could vary depending on the cell and tissue type and the conditions used to trigger autophagy, with cells deriving the membranes from the most suitable or expendable source. Thus, in a tissue with a defined function, in response to a specific stress stimulus, the growing autophagosome would be supplied with membranes from the most optimal reservoir: an organelle that could guarantee the delivery of a large amount of lipids, but whose depletion ideally would not adversely affect the function of the tissue. From a cursory look at the current available data, one might conclude that fasting animals utilize ER while nitrogen-starved yeast use Golgi, for example. However, accurate comparative studies are needed to determine whether such trends hold true for a wider variety of cell types and stress conditions.
Another possibility that should not be discarded a priori is that autophagosomes could be a mosaic of membranes derived from more than one organelle. For example, the phagophore could originate from one organelle, while additional lipid bilayers required for its expansion are acquired from another source. Having a spectrum of membrane sources to choose from could help ensure the availability of a large supply of lipids to sustain the progression of autophagy. One could imagine that a single intracellular organelle could not provide enough lipids to produce the multitude of autophagosomes generated during prolonged periods of starvation or stress.
Finally, it still remains to be determined whether the different organelles implicated so far in autophagosome biogenesis contribute to nonselective bulk autophagy or to selective forms of autophagy, such as mitophagy (selective degradation of mitochondria), pexophagy (selective degradation of peroxisomes), or reticulophagy (selective degradation of the ER). In this regard, the observation of ER or mitochondria connected to membranes of the phagophore, for example, could suggest that rather than contributing to autophagosome formation, these organelles are actually the object of the degradation process.
Autophagosomal membranes and drug therapies
Experimental evidence indicates that autophagosome biogenesis is probably a very complex process on several levels, including its regulation in response to different cellular and environmental cues, and the factors governing the choice of membrane sources.
Is there any therapeutic value in determining the origin of the autophagosomal membranes? We think that elucidating this process could ultimately provide new drug targets for the treatment of diseases that can be alleviated or cured by the activation of autophagy, including specific muscular dystrophies, persistent infections, and neurodegenerative disorders (ataxias, Huntington’s, and Parkinson’s diseases). Understanding the sources and processes by which the autophagosome’s lipid bilayers are delivered will undoubtedly reveal critical new proteins and articulate their functions, allowing researchers to pinpoint specific parts of the pathway.
Importantly, autophagy has also been associated with cancer. For example, loss of one of the two copies of Beclin1/Atg6, a gene involved in autophagy, is often found in human breast, ovarian, and prostate cancers.12 Similarly, alterations of factors regulating the trafficking of the transmembrane protein Atg9 have been found to be a direct cause of tumorigenesis. These observations support the possibility that specific illnesses could be the phenomenological manifestation of a misregulation of lipid bilayer flux during autophagy. As a result, the factors modulating these pathways would be optimal targets for drugs aimed at restoring normal membrane supply and consequently proper progression of autophagy.
Conclusions and future directions
Since the discovery of autophagy, the membrane origin of the autophagosomes has been the subject of intense debate. Recent studies employing advanced technologies have confirmed and extended the pioneering ultrastructural observations and have provided some insights on the membrane origin of these unique vesicles. The diverse conclusions of the recent work, however, have not yet provided an unequivocal answer, but rather have raised new questions that now need to be addressed. The work on this topic has only just begun.
Muriel Mari and Fulvio Reggiori are researchers in the Department of Cell Biology at the Utrecht University Medical Center, the Netherlands. Sharon A. Tooze is an investigator at the Cancer Research UK London Research Institute.
This article is adapted from a review in F1000 Biology Reports, DOI:10.3410/B3-25.
References
1. M. Hayashi-Nishino et al., “A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation,” Nat Cell Biol, 11:1433-37, 2009.
2. P. Ylä-Anttila et al., “3D tomography reveals connections between the phagophore and endoplasmic reticulum,” Autophagy, 5:1180-85, 2009.
3. E.L. Axe et al., “Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum,” J Cell Biol, 182:685-701, 2008.
4. K. Matsunaga et al., “Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L,” J Cell Biol, 190:511-21, 2010.
5. D.W. Hailey et al., “Mitochondria supply membranes for autophagosome biogenesis during starvation,” Cell, 141:656-67, 2010.
6. I.Tanida et al., “LC3 conjugation system in mammalian autophagy,” Int J Biochem Cell Biol, 36:2503-18, 2004.
7. Z. Xie, D.J. Klionsky, “Autophagosome formation: core machinery and adaptations,” Nat Cell Biol, 9:1102-09, 2007.
8. T. Yoshimori, T. Noda, “Toward unraveling membrane biogenesis in mammalian autophagy,” Curr Opin Cell Biol, 20:401-07, 2008.
9. J. Huang et al., “Antibacterial autophagy occurs at PI(3)P-enriched domains of the endoplasmic reticulum and requires Rab1 GTPase,” Autophagy, 7:17-26, 2011.
10. B. Ravikumar et al., “Plasma membrane contributes to the formation of pre-autophagosomal structures,” Nat Cell Biol, 12:747-57, 2010.
11. M. Mari et al., “An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis,” J Cell Biol, 190:1005-22, 2010.
12. S. Jin, E. White, “Role of autophagy in cancer: management of metabolic stress,” Autophagy, 3:28-31, 2007
Interested in reading more?





