Ancestral sequence reconstruction relies on phylogeny and statistics to infer the most likely amino acid
SEQUENCE ALIGNMENT
Scientists collect sequences from databanks of the modern versions of the protein of interest from different organisms.

TREE BUILDING
Computer algorithms construct a phylogenetic tree for the proteins (Curr Opin Struct Biol, 38:37–43, 2016).
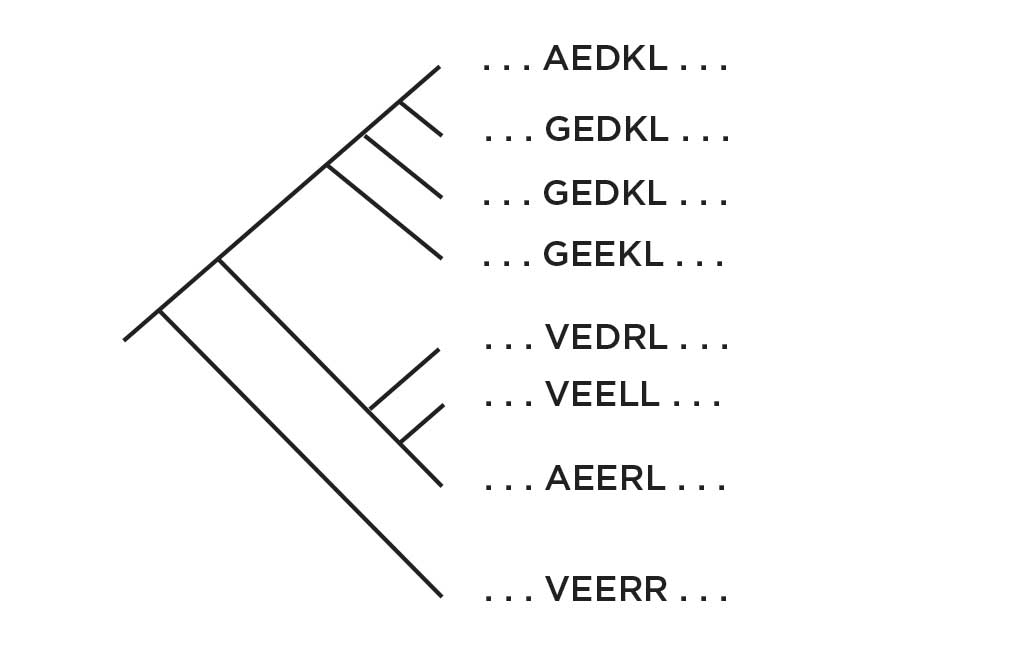
ANCESTRAL RECONSTRUCTION
The programs can then infer the sequences that likely existed at nodes of the tree, before the modern species evolved.

LABORATORY TESTS
Finally, the scientists order synthetic DNA and generate those proteins in the lab to use for experiments.

Ensuring accuracy
One way to ensure that an ASR protein behaves like the true ancestor is to resurrect and test not only the best amino acid sequence generated by the algorithms, but a few proteins with the second-best guesses, or third-best guesses, and so on. If those alternative ancestors act...

The Perfect Starting Point
Bioengineers love resurrected proteins because they often combine two desirable features: thermostability and promiscuity. For example, researchers at the University of Granada in Spain reconstructed several versions of an antibiotic-resistance protein called beta-lactamase, going back as far as 3 million years. As the protein evolved, its melting point dropped from more than 80 °C to less than 60 °C. It also became more specific for penicillin, losing its ability to neutralize other drugs (J Am Chem Soc, 135:2899–902, 2013).
Read the full story.
Interested in reading more?






