 UC DAVIS, GORILLA DOCTORS
UC DAVIS, GORILLA DOCTORS
Inbreeding apes
Species: Mountain gorilla (Gorilla beringei beringei)
The first full genome sequence of the mountain gorilla, one subspecies of the eastern gorilla, suggests a that the population has been declining for 100,000 years, resulting in limited genetic diversity and the highest known rate of primate inbreeding, according to a study published last week (April 9) in Science.
An international team of scientists sequenced the genomes of seven mountain gorillas and six eastern lowland gorillas (Gorilla beringei graueri), another eastern gorilla subspecies, comparing them to that of another gorilla species, the western gorilla (Gorilla gorilla), which split from the eastern gorilla 20,000 years ago. The researchers found that while 13.8 percent of a western lowland gorilla’s genome contained identical alleles on both sets of chromosomes, the same was true of 34.5 percent of the mountain gorilla genome and...
“I had thought beforehand that it was probably modern human influence in the last centuries that had really decimated the mountain gorillas,” study coauthor Chris Tyler-Smith of the Wellcome Trust Sanger Institute in the U.K. told The Verge, referring to the two remaining isolated populations of mountain gorilla on the borders of Rwanda, Uganda, and the Democratic Republic of Congo, which together number approximately 880 individuals. Instead, Tyler-Smith added, the study revealed that the population “has been small for thousands of years, so there’s no reason it shouldn’t continue this way for thousands more.”
 CDC, JANICE HANEY CARR
CDC, JANICE HANEY CARR
Fighting CF
Species: Pseudomonas aeruginosa
Genome size: 6.3 million base pairs
Sequencing the essential genome of a bacterium that sickens cystic fibrosis (CF) patients by growing into large, antibiotic-resistant colonies in has helped scientists to develop and refine a synthetic mix of chemicals that closely approximates the lung mucus of these patients, according to a study published last month (March 16) in PNAS.
The full genome of Pseudomonas aeruginosa was first reported 15 years ago, but researchers lack a good laboratory model to study the genes that make the bacterium so efficient at spreading in CF patients’ lungs. To overcome this problem, scientists from the University of Texas at Austin grew approximately 100,000 mutants from two different P. aeruginosa strains in normal laboratory media and in mucus samples from patients with CF to determine which genes were required for the bacterium’s survival. They found that the two strains had slightly different “essential genomes,” or sets of genes that were required, and that the essential genome also differed between lab media and CF patient-derived mucus, which was comparatively nutrient-poor. Next, the researchers used the bacteria to test the similarity of synthetic mucus to patient samples by determining the essential genomes of the bugs in each environment. They found that only two gene mutants of both strains were significantly less fit in the synthetic mucus than they were in the patient sample. Other strain-specific mutants that behaved differently in synthetic mucus compared to patient samples could be rescued by adding the products from a limited number of biochemical pathways to the synthetic liquid.
“We’ve developed something other labs can replicate,” said study coauthor Marvin Whiteley of the University of Texas in a statement, referring to the synthetic mucus. “It allows researchers to do relevant experiments in a context that really matters.”
 MPI F. ORNITHOLOGY, S. SELTMANN
MPI F. ORNITHOLOGY, S. SELTMANN
Musical hormones
Species: Domesticated canary (Serinus canaria)
Genome size: 1.2 billion base pairs
The full genome of a female canary (Serinus canaria) and brain gene expression patterns of male conspecifics has revealed a unique set of hormone-regulated genes involved in birds’ complex songs, according to a study published in Genome Biology earlier this year (January 29).
Originally domesticated from wild birds on the Canary Islands in the 15th century, canaries change their singing patterns based on the season and on hormone levels, thus serving as a useful model of brain plasticity. To explore the genetic basis of canary song, researchers from the Max Planck Institutes for Ornithology and Molecular Genetics, both in Germany, sequenced the genome of a female canary. Aligning the sequence to that of a diverse set of 13 other birds revealed a similar genome structure among the different species.
To search for canary-specific genes involved in birdsong, the scientists then recorded and analyzed the music of male canaries, some of whom had testosterone implants to induce breeding-season stereotypical songs while the others sang variable non-breeding season tunes. The researchers then sequenced the expressed genes from two brain regions known to be involved in hormone-dependent singing and compared them to the genes expressed by zebra finches, whose songs are not influenced by hormones. In one of the brain regions, the scientists found that 35 percent of the canary genes that responded to estrogen and 11 percent that responded to androgen were hormone-independent in zebra finches, meaning that these variants were canary-specific.
“In the canary, those genes that are sensitive for testosterone and estrogen, and also involved in the rewiring of neurons, might be crucial for . . . seasonal song patterning,” study coauthor Stefan Leitner of the Max Planck Institute for Ornithology wrote at the BioMed Central blog On Biology.
 CDC, ERSKINE PALMER
CDC, ERSKINE PALMER
Mixed-up bird flu
Strain: H5 clade 2.3.4.4 reassortant H5N1 avian influenza virus
A strain of bird flu detected in a green-winged teal shot in Washington State on December 29, 2014, provides evidence of yet another new mixture of H5N8, a highly pathogenic Eurasian virus, and low pathogenic influenza circulating in wild North American birds, according to a study published this month (April 2) in Genome Announcements.
The H5N8 virus, which has been detected in birds in South Korea, Japan, Russia, and Europe, arrived in North America last year, and had already combined with North American strains to make a H5N2 virus that has infected commercial and wild flocks of many bird species across the United States.
Scientists from the US Geological Survey (USGS) and the US Department of Agriculture sequenced the virus from the teal and found that it represents a separate mixture of Eurasian H5N8 and low pathogenic North American flu strains. The genome reveals that it is not the same H5N1 detected in Asia in 1996.
“This report describes the first detection of [highly pathogenic avian influenza] H5N1 virus in North America, and this virus has since been detected in a backyard flock in British Columbia, Canada,” study coauthor Hon Ip of the USGS National Wildlife Health Center scientist said in a statement.
Pathogenicity is measured by the ability of the strain to produce disease in chickens, so it is not known how the new strain will impact wild birds. There have been no human infections and the public health risk is considered low.
 CDC, PUBLIC HEALTH IMAGE LIBRARY
CDC, PUBLIC HEALTH IMAGE LIBRARY
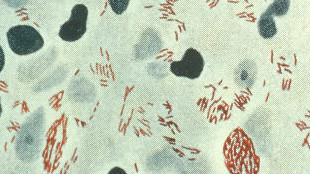
Leprosy bacterium
Species: Mycobacterium lepromatosis
Genome size: 3.27 million base pairs
The recently sequenced genome of a species of bacterium thought to cause a rare and more severe form of leprosy than the traditional Mycobacterium leprae resembles but differs from the previously sequenced M. leprae, according to a study published last month (March 23) in PNAS.
M. leprae, which was discovered as the cause of leprosy in the late 19th century, had already been sequenced from modern and medieval samples in 2013. In the latest study, researchers from the Ecole Polytechnique Fédérale in Lausanne, Switzerland, sequenced a second species of bacteria, M. lepromatosis, which was first reported in Mexico and Caribbean in 2008 and is thought to cause diffuse lepromatous leprosy.
Comparing the two leprosy bacteria genomes revealed strong similarities in both protein-coding genes (93 percent identity) and pseudogenes (82 percent identity), although the scientists estimated that the species diverged 13.9 million years ago. Both modern species and their common ancestor display evidence of heavy gene loss and decay, making it impossible for the bacteria to survive without a host to provide nutrients and suggesting that their common ancestor also caused leprosy. The species differed somewhat in which genes they were missing, although testing the significance of this observation will be difficult. “We cannot grow these bacteria in the lab, so we cannot conduct genetic experiments,” study coauthor Andrej Benjak told The Scientist.
Skin samples from Asia, Africa, and the Americas suggested that while the better-known M. leprae is widespread, M. lepromatosis appears to be confined to Mexico. The bacterium has genes that would allow it to invade Schwann cells in the nervous system, like M. leprae, but it is unknown if this occurs.
Interested in reading more?




