Scientists are learning that epigenetic marks such as DNA methylation can influence not just gene expression, but genes themselves. When this happens in germline cells, the changes can diversify the genetic substrate upon which natural selection acts. Scientists are still exploring how such processes could influence evolutionary processes overall.
Triggering mutations

Scientists have known for several decades that methylation can cause the nucleobase cytosine to lose an amino group, transforming it into thymine. When this occurs during gametogenesis, it can be one of the major drivers of evolution at the molecular scale. Some human embryo studies estimate that for particular genomic regions, some 20 percent of mutations unique to humans arose from cytosine methylation.
Shaping mutation patterns

Some studies in cancer cells suggest that the packaging of DNA into dense structures called nucleosomes not only shapes the expression of those genes, but also their mutation rates. When bound in such conformations, DNA is less likely to open into a single-stranded state that makes it more prone to mutations.
Dampening mutation
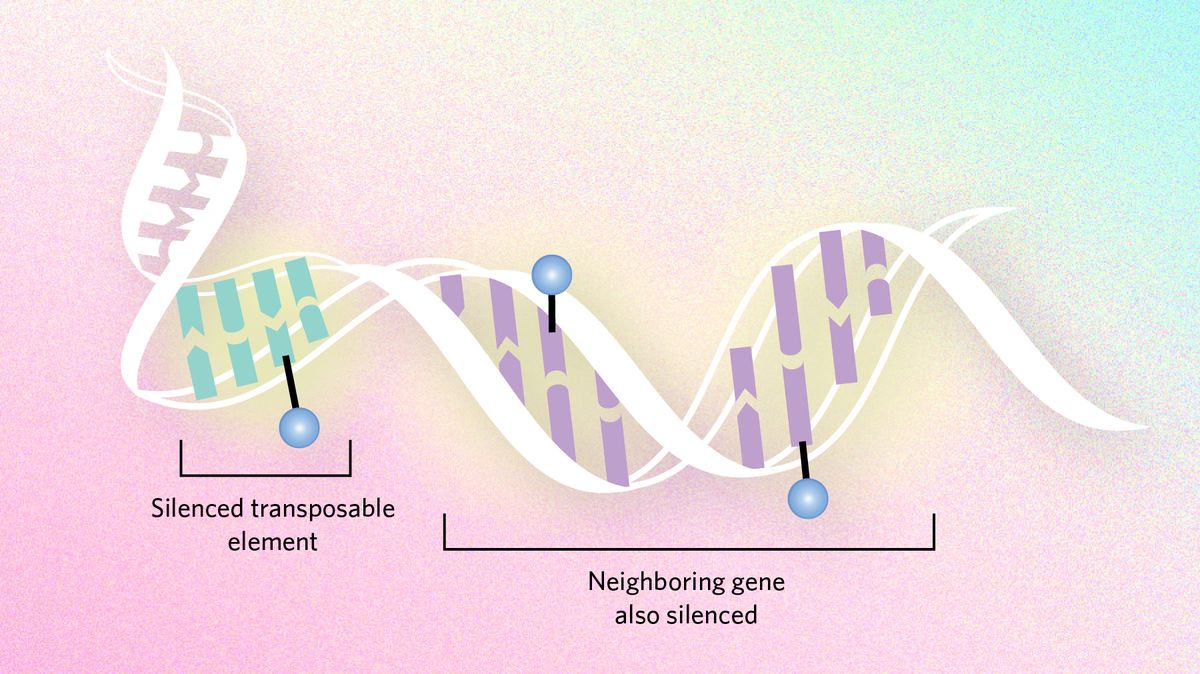
Epigenetic dynamics can also slow evolutionary processes by suppressing transposable elements, snippets of DNA that hop around the genome and cause mutations. Genes neighboring these troublemakers can end up being silenced too, sometimes creating a conflict between the need to express genes and the need to silence transposons. In certain Drosophila species, researchers hypothesize that the flies have moved large chunks of the transposon-infested Y chromosome to other chromosomes to ensure important genes are not silenced.
New gene functions

Long-term epigenetic silencing of particular genes could allow them to accumulate genetic mutations and develop new functions over time. There is some correlational evidence that duplicated genes—which are often lost from the genome unless they are beneficial to the organism—frequently become epigenetically silenced. They could thereby evolve new functions.
Read the full story.
