Following chemist and marine biologist Osamu Shimomura’s success in determining the structure of luciferin from a crustacean at Nagoya University, Frank Johnson, then a biologist at Princeton University, invited him to study luminescence in his lab. When Shimomura arrived at Princeton University in 1960, he had no ambitions for novel proteins and their applications.
Johnson was interested in in the blue light produced by a species of jellyfish called Aequorea aequorea (now known as Aequorea victoria), Shimomura began looking into the phenomenon. At the time, scientists believed that all luminescence involved luciferin and luciferase. However, the duo did not purify any luciferin-related proteins from the jellyfish. This led Shimomura to believe that the blue light must come from another protein, but he struggled to isolate it.
A new way to luminesce
Shimomura attempted to isolate this mystery protein under various conditions. He finally found that he could inactivate the luminescence with acid and obtain cell-free extract from the jellyfish tissues. Satisfied with this finding, he dumped the used samples down the sink. To his surprise, they glowed bright blue. He reasoned that the substrate for his mystery luminescent protein must be in the sink, and that it certainly wasn’t luciferase. The aquarium in the lab drained into the sink, so Shimomura got to work testing the components in it to see if they elicited blue light from his samples. Calcium turned out to be the culprit.
By removing calcium from the samples, Shimomura and Johnson isolated the protein responsible for the blue light, which they named aequorin.1 This was the first example of a light-activated protein, which they called a photoprotein.
I wanted to have a tool like this. I thought this would be wonderful.
—Martin Chalfie, Columbia University
Today, we call them fluorescent proteins. Alongside aequorin, Shimomura and Johnson also purified a small amount of another protein that glowed green instead of blue. They called it “green protein,” but at the time, they could not purify sufficient amounts to study it further. While collecting and saving these samples for future analysis, they continued exploring aequorin to characterize its structure and chemistry.
Across the country in the mid-1960s, James Morin, a field biologist who is now a professor emeritus at Cornell University, became fascinated by bioluminescence. “It was amazing to me that so many organisms that didn’t really have the capacities for detecting luminescence would be luminescent,” he said.
This curiosity led him to the hydrozoan Obelia geniculata when he began his graduate studies at Harvard University. He noticed that the luminescence he saw from these animals in the ocean was green, whereas in his prepared samples, it was blue. “That indicated there must be some kind of coupling,” Morin said.
It turned out that O. geniculata, like Shimomura’s and Johnson’s A. aequorea, used a calcium-activated protein that emitted blue light. This light activated a second protein that glowed green. Morin published his findings from O. geniculata and coined the term “green fluorescent protein” (GFP) for the first time in 1971.2 Shimomura confirmed that the same events happened in his jellyfish in 1974.3
Shimomura continued to study GFP and identified its chromophore in 1979.4 Ultimately, though, he returned to studying bioluminescence. Morin also was more interested in the function of luminescence and the nervous system of the simple marine species that produced it. “We were just interested in the mechanisms of how that whole system worked,” Morin said. “I’ve never been an applications biologist. I’ve been a discovery kind of biologist.”
For the next two decades, GFP remained just an interesting luminescent protein found in some species of marine animals.
Not just a jellyfish thing
In the 1970s, Morin, then at the University of California, Los Angeles, took Paul Brehm on as a graduate student and introduced him to Obelia and the field of bioluminescence. As a biologist at Oregon Health and Science University, Brehm continued studying bioluminescence in Obelia as an associate professor. In 1989, he presented his work at a seminar at Columbia University; Martin Chalfie, a geneticist at Columbia University who had never heard of GFP before, happened to attend. After Brehm’s talk, Chalfie started tracking down GFP scientists on the phone. “I was looking for where genes were expressed,” Chalfie said. “I wanted to have a tool like this. I thought this would be wonderful.”
In 1989, scientists studied genes of interest in cells using newly developed technologies. “But in all of these instances, whether it’s antibodies, or in situ hybridization, or using a reporter like beta galactosidase, one needed to really prepare the samples. That meant first they were fixed,” Chalfie said. “As a result, one had really a snapshot in time.” There was no way to visualize what genes were active in a living organism.
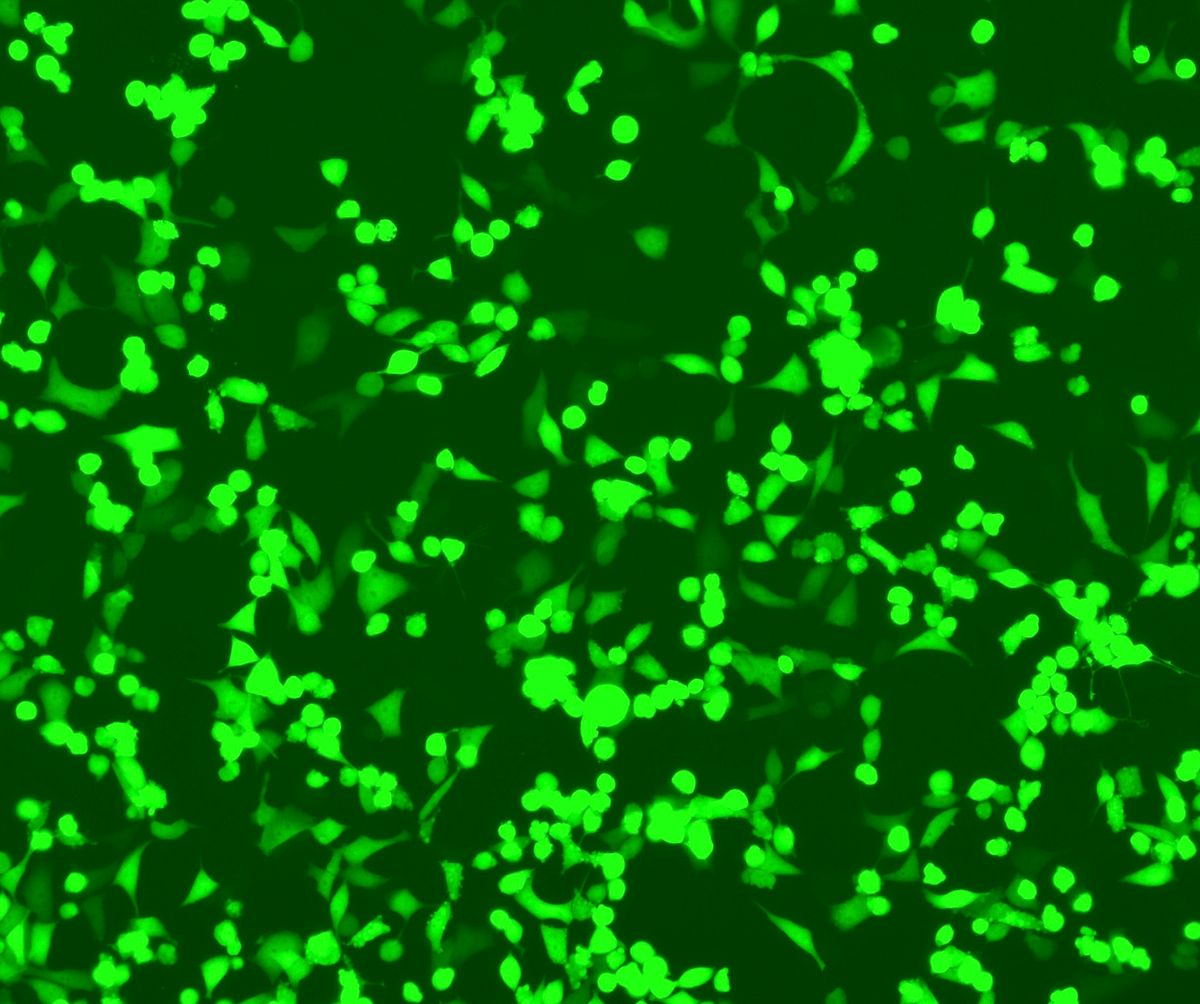
Chalfie thought that if GFP could be taken out of these marine organisms and inserted into other cells, it could act as a reporter in a living animal. However, nobody knew if this would be possible. GFP had a cyclized portion in its structure, and many people thought that such a complex chromophore structure would need the help of other enzymes in the jellyfish to fold correctly.
Chalfie’s phone calls eventually led him to Douglas Prasher, a molecular biologist who worked at the Woods Hole Oceanographic Institute. Prasher cloned the aequorin genes in 1985 and began working on cloning the gene for GFP.5 The two struck up a collaboration. In 1992, Prasher published his paper describing the GFP cDNA clone, which he sent to Chalfie and his graduate student, Ghia Euskirchen.6
With the clone in hand, the two had a decision to make. Since Prasher cut the GFP gene out of the jellyfish genome with restriction digestion enzymes, the clone had extra DNA segments that bookended the GFP gene. If Chalfie and Euskirchen wanted only the GFP DNA, they could amplify just the GFP gene by PCR, but that risked introducing mutations since PCR was error-prone at the time. Alternatively, they could transform the whole clone into E. coli, which would generate high-fidelity DNA but retain the extra jellyfish sequence segments. Because of this risk, most groups opted for transforming the whole clone. “I know at least three groups that did that, and it did not work,” Chalfie said. “We did something different.”
Taking a risk on error-prone PCR, Euskirchen amplified only the GFP coding sequence and inserted it into an expression vector to transform back into E. coli. When Euskirchen imaged her bacteria using a fluorescent microscope, she saw green glowing back at her.
“That immediately said there is nothing else needed from the jellyfish,” Chalfie said.
While this was a huge milestone in molecular biology, Chalfie believes that he owes some of his success to other contributing factors. “It’s not that I was the first person in the world who ever thought about this,” Chalfie said. “It was a combination of being at the right place at the right time, having the right people in the lab and the right expertise to be able to try it, and then a smidgen of luck.”
With this success, Chalfie and his team tested the initial goal of inserting the isolated GFP into the touch-sensing cells of Caenorhabditis elegans. With positive data in hand, the group published the first manuscript with GFP as a reporter in a nonmarine animal.7
An elegant measurement
Once the word was out that GFP could be put into other model organisms, the protein found itself visiting labs across the country, quite literally in the case of Raphael Valdivia, who is currently a molecular geneticist at Duke University and who was a graduate student in Stanley Falkow’s lab at Stanford University in the mid-1990s.
“This was when I had to take my qualifying exams to get into PhD candidacy, and so I had to propose a project outside of my thesis project,” said Valdivia. “I proposed to use GFP as a reporter for gene expression in bacteria.”
How do you make a really elegant measurement that’s biologically relevant and learn things with it?
—Geoffrey Baird, University of Washington
Valdivia got the clone from Prasher and amplified the GFP coding sequence to reinsert it into an expression vector, similar to Chalfie’s experiments, and then transformed it into E. coli. “But when we expressed it, it did not work very well,” Valdivia explained. Although the bacteria were green, the GFP took a long time to fold and become fluorescent. “The jellyfish version of GFP just crashed out in E. coli,” he said.
Valdivia wondered if he could tweak GFP to brighten its fluorescence. Working alongside Brendan Cormack, he created mutants of GFP that targeted the protein’s chromophore.8 They had their brighter mutants within a week.
“We didn’t do it with an idea of doing big technological breakthroughs or screens in general. We just needed something that worked better in pathogens so we could screen,” Valdivia said.
Valdivia eventually gave GFP to the spouse of a postdoctoral fellow in Falkow’s lab who worked at ClonTech, which is now Takara Bio USA, Inc. Through this spurious connection, ClonTech optimized GFP for expression in mammalian cells, licensed, and commercialized as enhanced GFP, or eGFP in 1996.
While many groups worked with GFP to change its stability or activity, Roger Tsien, a chemist at the University of California, San Diego made possibly the most striking contributions. With a background in dye chemistry, Tsien saw the value a fluorescent protein could offer biological research. According to Geoffrey Baird, who was a graduate student in Tsien’s lab and is now a clinical pathologist at the University of Washington, his mentor had a fundamental interest that drove his research. “How do you make a really elegant measurement that’s biologically relevant and learn things with it?” Baird asked.
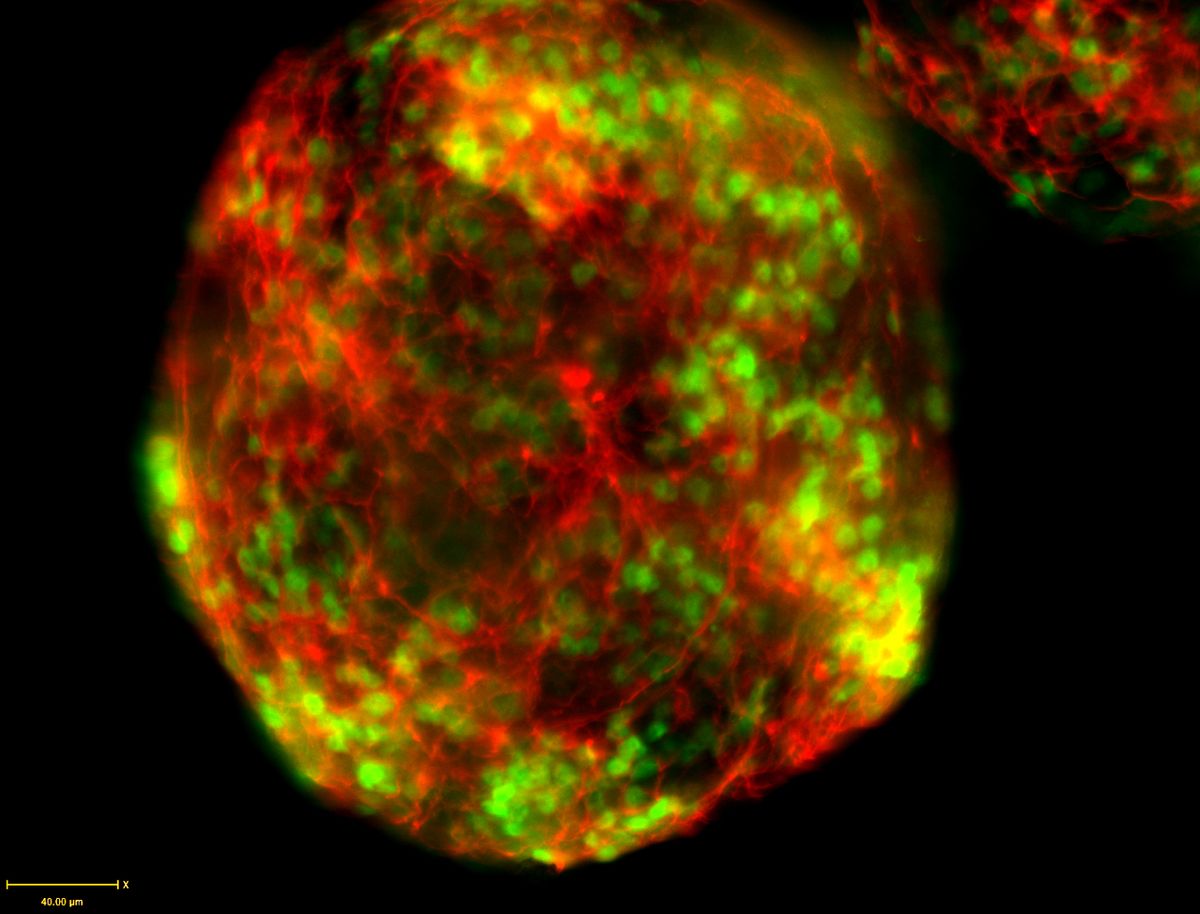
Tsien’s goal was to use GFP in fluorescence resonance energy transfer (FRET) to monitor calcium signaling.9 Although his team successfully generated a few mutants that gave rise to altered colors of GFP, Tsien anticipated that solving the structure of GFP would be important for more modifications.10,11
“Because it was only really with the structure that one could then sort of say, well, hey, if we made this mutation here, we might be able to make it a much different color,” Baird explained.
Tsien’s group solved this structure with help from James Remington’s group at the University of Oregon in 1996.12 Within a week, a second group published their finding of the structure, and the two closely aligned.13 With this information, Tsien’s group got to work exploring what was possible with GFP. “I basically got into the business of making mutations,” Baird said.
Baird considers his most important contribution to have come from an accident. He was trying to create brighter GFP clones, and one day, he saw one that didn’t get dimmer after acid exposure. After he sequenced it, Baird realized that he had made a mistake. “This thing shouldn’t be fluorescent at all,” he said.
Instead of a single point mutation, he had accidentally inserted six amino acids into the sequence. But this accident showed that it was possible to distort GFP. Baird informed Tsien of his mistake. “He basically had all of the ideas for what you would do to exploit a serendipitous finding,” Baird said.
Ultimately, the team found that breaking GFP and rejoining it in a variety of ways through a process called circular permutation created proteins with applications not only in FRET, but also for studying protein interactions just by observing if a split GFP molecule became active again.14 As Tsien had hoped and anticipated, GFP was a very useful measuring stick.
Revolutionizing biology
From an obscure protein that wasn’t even abundant enough to study, scientists have now published more than 40,000 papers that reference green fluorescent protein. Today, GFP tags proteins to view them inside of cells under microscopes and to isolate cells of interest using flow cytometry and fluorescence-activated cell sorting.15 It has confirmed physical connections between proteins and determined if a protein is made in the first place. GFP has found itself in viruses, yeast, plants, mice, rats, rabbits, pigs, and even nonhuman primates.16-23
“It revolutionized cell biology and molecular biology,” said Baird.
Shimomura, Chalfie, and Tsien received the Nobel Prize in 2008 for their work on GFP.
“It’s been useful in the way I thought,” Chalfie said, reflecting on GFP’s legacy. “But even more so, it’s been useful in ways that I would have never imagined.”
- Shimomura O, et al. Extraction, purification, and properties of Aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Physiol. 1962;59(3):223-239
- Morin JG & Hastings JW. Energy transfer in a bioluminescent system. J Cell Physiol. 1971;77(3):313-318
- Morise H, et al. Intermolecular energy transfer in the bioluminescent system of Aequorea. Biochemistry. 1974;13(12):2656-2662
- Shimomura O. Structure of the chromophore of Aequorea green fluorescent protein. FEBS Lett. 1979;104(2):220-222
- Prasher D, et al. Cloning and expression of the cDNA coding for aequorin, a bioluminescent calcium-binding protein. Biochem Biophys Res Commun. 1985;126(3):1259-1268
- Prasher DC, et al. Primary structure of the Aequorea Victoria green-fluorescent protein. Gene. 1992;111(2):229-233
- Chalfie M, et al. Green fluorescent protein as a marker for gene expression. Science. 1994;263(5148):802-805
- Cormack BP, et al. FACS-optimized mutants of the green fluorescent protein (GFP). Gene. 1996;173(1)33-38
- Sekar RB & Periasamy A. Fluorescence resonance energy transfer (FRET) microscopy imaging of live cell protein localizations. J Cell Biol. 2003; 160(5):629-633
- Heim R & Tsien RY. Engineering green fluorescent protein for improved brightness, longer wavelengths, and fluorescent resonance energy transfer. Curr Biol. 1996;6(2):178-182
- Heim R, et al. Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc Natl Acad Sci. 1994;91(26):12501-12504
- Ormö M, et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science. 1996;273(5280):1392-1395
- Yang F, et al. The molecular structure of green fluorescent protein. Nat Biotechnol. 1996;14:1246-1251
- Baird GS, et al. Circular permutation and receptor insertion within green fluorescent proteins. Proc Natl Acad Sci. 1999;96(20):11241-11246
- Chudakov DM, et al. Fluorescent Proteins and Their Applications in Imaging Living Cells and Tissues. Physiol Rev. 2010;90(3):1103-1163
- Oparka KJ, et al. Using GFP to study virus invasion and spread in plant tissues. Nature. 1997;388:401-402
- Li J, et al. Green fluorescent protein in Saccharomyces cerevisiae: real-time studies of the GAL1 promoter. Biotechnol Bioeng. 2000;70(2):187-196
- Köhler RH. GFP for in vivo imaging of subcellular structures in plant cells. Trends Plant Sci. 1998;3(8):317-320
- Yang M, et al. Visualizing gene expression by whole-body fluorescence imaging. Proc Natl Acad Sci. 2000;97(22)12278-12282
- Hakamata Y, et al. Green fluorescent protein-transgenic rat: a tool for organ transplantation research. Biochem Biophy Res Commun. 2001;286(4):779-785
- Takahashi R, et al. Establishment and characterization of CAG/EGFP transgenic rabbit line. Transgenic Res. 2007;16:115-120
- Brunetti D, et al. Transgene expression of green fluorescent protein and germ line transmission in cloned pigs derived from in vitro transfected adult fibroblasts. Cloning Stem Cells. 2008;10(4):409-420
- Kurre P, et al. Kinetics of fluorescence expression in nonhuman primates transplanted with GFP retrovirus-modified CD34 cells. Mol Ther. 2002;6(1):83-90















