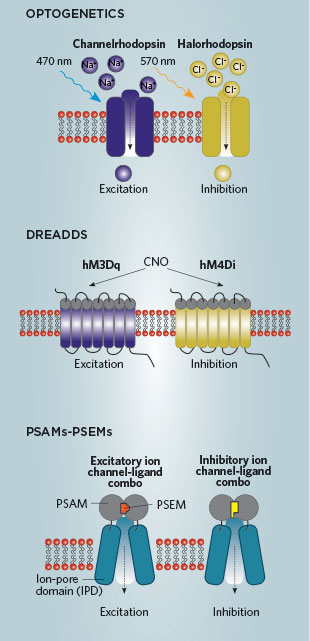
 A MATTER OF TIME: Optogenetics methods, which work on the millisecond timescale, allow for the finest level of temporal control over neuron excitation and inhibition. The chemogenetic tools, DREADDs and PSAMs-PSEMs, are ideal for the study of longer-lasting behaviors such as appetite, thirst, or anxiety because they work over a scale of the minutes-to-hours. The receptors are incorporated into specific neurons or cells using viruses. Ligands—CNO or salvinorin B (for DREADD receptors) or PSEMs—are administered via injection or drinking water. Both receptors and ligands are orthogonal, meaning they do not bind to anything else in the body.REDRAWN WITH PERMISSION OF SCOTT STERNSON FROM SCIENCE, 333:1292, 2011; NEUROENDOCRINOLOGY, 100:98, 2014In a pharmacology lab at the University of North Carolina at Chapel Hill, doctoral student Reid Olsen, working with brain tissue harvested from a mouse just a few hours earlier, readies half a dozen dime-size slices for live calcium imaging. This mouse’s brain contains a genetically engineered receptor that Olsen has targeted to cells thought to control the making of new neurons in adult mice. He is about to use a synthetic drug to activate this receptor in the tissue. When it indeed works—just as he has predicted—he turns his attention to attempting to stimulate neurogenesis in a freely moving mouse that has the same engineered receptors in its brain.
A MATTER OF TIME: Optogenetics methods, which work on the millisecond timescale, allow for the finest level of temporal control over neuron excitation and inhibition. The chemogenetic tools, DREADDs and PSAMs-PSEMs, are ideal for the study of longer-lasting behaviors such as appetite, thirst, or anxiety because they work over a scale of the minutes-to-hours. The receptors are incorporated into specific neurons or cells using viruses. Ligands—CNO or salvinorin B (for DREADD receptors) or PSEMs—are administered via injection or drinking water. Both receptors and ligands are orthogonal, meaning they do not bind to anything else in the body.REDRAWN WITH PERMISSION OF SCOTT STERNSON FROM SCIENCE, 333:1292, 2011; NEUROENDOCRINOLOGY, 100:98, 2014In a pharmacology lab at the University of North Carolina at Chapel Hill, doctoral student Reid Olsen, working with brain tissue harvested from a mouse just a few hours earlier, readies half a dozen dime-size slices for live calcium imaging. This mouse’s brain contains a genetically engineered receptor that Olsen has targeted to cells thought to control the making of new neurons in adult mice. He is about to use a synthetic drug to activate this receptor in the tissue. When it indeed works—just as he has predicted—he turns his attention to attempting to stimulate neurogenesis in a freely moving mouse that has the same engineered receptors in its brain.
Less than a decade ago, such precise control over neuronal activity in a dish, let alone in a living brain, was impossible. The drugs available to repress neurons or encourage them to fire would produce off-target effects or eliminate cell populations indiscriminately.
Working in the lab of Juan Song, Olsen is using a “designer receptor exclusively activated by a designer drug,” or DREADD. These modified G protein–coupled receptors (GPCRs) are usually either virally administered or bred into animals, then activated by a specific ligand that’s either injected or taken orally. Both the receptor and the ligand are designed to be orthogonal, effectively meaning they bind to each other but to nothing else.
Along with DREADDs, recently developed orthogonal ligand-gated ion channels called “pharmacologically selective actuator ...